Multiple Variables
- Level: 🟢 Beginner
- Est. Time: 15 minutes
- Concepts: Multiple Variables, Variable Dependencies, Coupled Mechanisms, System Dynamics
This example provides:
- Complete multi-variable simulation with coupled dynamics
- Three competing ecological theories
- System-level governance for ecosystem stability
- Detailed analysis including correlation and cycle detection
- Phase portrait visualization
- Cross-correlation analysis
- Exercises for extension
Overview
This example demonstrates how to work with multiple interconnected variables in Procela. We'll build a classic predator-prey ecosystem (Lotka-Volterra model) where prey (rabbits) and predator (foxes) populations interact.
The simulation features:
- Two variables (Rabbits and Foxes) with mutual dependencies
- Multiple mechanisms proposing different ecological theories
- Variable coupling where changes in one affect the other
- Governance monitoring ecosystem stability
from typing import Any
import matplotlib.pyplot as plt
import numpy as np
from matplotlib.gridspec import GridSpec
from scipy.signal import find_peaks
from procela import (
Executive,
InvariantPhase,
InvariantViolation,
Key,
Mechanism,
RangeDomain,
SystemInvariant,
Variable,
VariableRecord,
VariableSnapshot,
WeightedConfidencePolicy,
)
rng = np.random.default_rng(42)
INITIAL_SOURCE = Key()
Step 1: Create Variables
First, create variables for prey and predator populations:
# Prey population (Rabbits) - range 0 to 1000
rabbits = Variable(
name="Rabbits", domain=RangeDomain(0, 1000), policy=WeightedConfidencePolicy()
)
# Predator population (Foxes) - range 0 to 500
foxes = Variable(
name="Foxes", domain=RangeDomain(0, 500), policy=WeightedConfidencePolicy()
)
# Initialize populations
rabbits.init(VariableRecord(100.0, confidence=1.0, source=INITIAL_SOURCE))
foxes.init(VariableRecord(20.0, confidence=1.0, source=INITIAL_SOURCE))
print(f"Initial ecosystem: {rabbits.value:.0f} rabbits, {foxes.value:.0f} foxes")
Key Points: - Variables have different domains (rabbits can be more numerous than foxes) - Both start with 100% confidence initial values - Each variable has its own resolution policy
Step 2: Define Coupled Mechanisms
Mechanisms can read multiple variables and write to multiple variables:
Classic Lotka-Volterra Mechanism
class LotkaVolterraMechanism(Mechanism):
"""
Lotka volterra mechanism.
Classic predator-prey model:
- Prey growth: dr/dt = α*r - β*r*f
- Predator growth: df/dt = δ*r*f - γ*f
where:
α = prey birth rate
β = predation rate
δ = predator efficiency (how well predators convert prey to offspring)
γ = predator death rate
"""
def __init__(
self,
alpha: float = 0.08,
beta: float = 0.02,
delta: float = 0.01,
gamma: float = 0.06,
) -> None:
"""Lotka volterra mechanism constructor."""
super().__init__(reads=[rabbits, foxes], writes=[rabbits, foxes])
self.alpha = alpha # Prey birth rate
self.beta = beta # Predation rate
self.delta = delta # Predator efficiency
self.gamma = gamma # Predator death rate
def transform(self) -> None:
"""Transform method."""
# Read current populations
r = self.reads()[0].value # Rabbits
f = self.reads()[1].value # Foxes
# Calculate changes
dr_dt = r * (self.alpha - self.beta * f)
df_dt = f * (self.delta * r - self.gamma)
# Discrete time update (dt = 1)
new_rabbits = max(0, r + dr_dt)
new_foxes = max(0, f + df_dt)
# Add stochastic noise (environmental variability)
new_rabbits += rng.normal(0, new_rabbits * 0.05)
new_foxes += rng.normal(0, new_foxes * 0.05)
# Confidence decreases with extreme values
rabbit_confidence = 0.9 if 10 < new_rabbits < 500 else 0.6
fox_confidence = 0.9 if 5 < new_foxes < 200 else 0.6
# Propose hypotheses for both variables
self.writes()[0].add_hypothesis(
VariableRecord(
value=new_rabbits,
confidence=rabbit_confidence,
source=self.key(),
metadata={"model": "classic", "alpha": self.alpha},
)
)
self.writes()[1].add_hypothesis(
VariableRecord(
value=new_foxes,
confidence=fox_confidence,
source=self.key(),
metadata={"model": "classic", "gamma": self.gamma},
)
)
print(
f" 🦊 LV Model: R:{new_rabbits:.0f} (conf:{rabbit_confidence:.2f}), "
f"F:{new_foxes:.0f} (conf:{fox_confidence:.2f})"
)
Alternative: Logistic Prey with Predation
class LogisticPreyMechanism(Mechanism):
"""
Alternative model.
Prey growth is logistic (carrying capacity limited) rather than exponential.
"""
def __init__(
self,
carrying_capacity: float = 800,
predation_rate: float = 0.03,
predator_efficiency: float = 0.008,
predator_death: float = 0.07,
) -> None:
"""Logistic prey mechanism constructor."""
super().__init__(reads=[rabbits, foxes], writes=[rabbits, foxes])
self.K = carrying_capacity
self.beta = predation_rate
self.delta = predator_efficiency
self.gamma = predator_death
def transform(self) -> None:
"""Transform method."""
r = self.reads()[0].value
f = self.reads()[1].value
# Logistic prey growth: r * (1 - r/K)
prey_growth = r * (1 - r / self.K)
prey_death = self.beta * r * f
new_rabbits = max(0, r + prey_growth - prey_death)
# Predator growth (same as LV)
predator_growth = self.delta * r * f
predator_death = self.gamma * f
new_foxes = max(0, f + predator_growth - predator_death)
# Add noise
new_rabbits += rng.normal(0, new_rabbits * 0.03)
new_foxes += rng.normal(0, new_foxes * 0.03)
# Higher confidence when near equilibrium
equilibrium_r = self.K * (1 - self.gamma / (self.delta * self.K))
r_distance = abs(new_rabbits - equilibrium_r) / self.K
rabbit_confidence = max(0.5, 0.9 - r_distance)
self.writes()[0].add_hypothesis(
VariableRecord(
new_rabbits,
confidence=rabbit_confidence,
source=self.key(),
metadata={"model": "logistic"},
)
)
self.writes()[1].add_hypothesis(
VariableRecord(
new_foxes,
confidence=0.8,
source=self.key(),
metadata={"model": "logistic"},
)
)
print(
f" 🌿 Logistic Model: R:{new_rabbits:.0f} (conf:{rabbit_confidence:.2f}), "
f"F:{new_foxes:.0f}"
)
Simple Ratio Mechanism
class RatioMechanism(Mechanism):
"""
Simple model based on predator-prey ratio.
Assumes populations tend toward a fixed ratio.
"""
def __init__(self, target_ratio: float = 5.0, adjustment_rate: float = 0.1) -> None:
"""Ratio mechanism constructor."""
super().__init__(reads=[rabbits, foxes], writes=[rabbits, foxes])
self.target_ratio = target_ratio # Target rabbits:foxes ratio
self.rate = adjustment_rate
def transform(self) -> None:
"""Transform method."""
r = self.reads()[0].value
f = self.reads()[1].value
if f == 0:
new_foxes = 1.0 # Avoid division by zero
else:
current_ratio = r / f
# Adjust toward target ratio
ratio_error = current_ratio - self.target_ratio
adjustment = self.rate * ratio_error
# Apply adjustment
new_rabbits = max(0, r - adjustment * r)
new_foxes = max(0, f + adjustment * f)
# Simple model has lower confidence
confidence = 0.65
self.writes()[0].add_hypothesis(
VariableRecord(new_rabbits, confidence=confidence, source=self.key())
)
self.writes()[1].add_hypothesis(
VariableRecord(new_foxes, confidence=confidence, source=self.key())
)
print(
f" ⚖️ Ratio Model: R:{new_rabbits:.0f}, "
f"F:{new_foxes:.0f} (conf:{confidence:.2f})"
)
Step 3: Create Governance for Ecosystem Stability
class EcosystemStabilityGovernance(SystemInvariant):
"""Monitor ecosystem health and prevents extinction."""
def __init__(
self, rabbits_var: Variable, foxes_var: Variable, min_population: float = 10
) -> None:
"""Ecosystem stability governance constructor."""
self.rabbits = rabbits_var
self.foxes = foxes_var
self.min_population = min_population
self.interventions: list[dict[str, Any]] = []
def check(snapshot: VariableSnapshot) -> bool:
"""Check ecosystem health after each step."""
r = self.rabbits.value
f = self.foxes.value
if r < self.min_population or f < self.min_population:
print(f"\n 🚨 ECOSYSTEM CRISIS at step {snapshot.step}!")
print(f" Population too low: R={r:.0f}, F={f:.0f}")
# Intervention: Restore populations
if r < self.min_population:
print(
" Intervention: Restoring rabbit population "
f"to {self.min_population}"
)
if f < self.min_population:
print(
" Intervention: Restoring fox population "
f"to {self.min_population}"
)
self.interventions.append(
{"step": snapshot.step, "rabbits_before": r, "foxes_before": f}
)
return False
return True
def handle(invariant: InvariantViolation, snapshot: VariableSnapshot) -> None:
# Nothing to handle
pass
super().__init__(
name="EcosystemStability",
condition=check,
on_violation=handle,
phase=InvariantPhase.POST,
)
# We'll integrate this into the executive step
Step 4: Run the Multi-Variable Simulation
# Create all mechanisms
mechanisms = [
LotkaVolterraMechanism(alpha=0.08, beta=0.02, delta=0.01, gamma=0.06),
LogisticPreyMechanism(carrying_capacity=800, predation_rate=0.025),
RatioMechanism(target_ratio=4.0, adjustment_rate=0.08),
]
# Create stability governance
stability_gov = EcosystemStabilityGovernance(rabbits, foxes, min_population=10)
# Create executive
executive = Executive(mechanisms=mechanisms)
executive.add_invariant(stability_gov)
print("\n" + "=" * 60)
print("Predator-Prey Ecosystem Simulation")
print("=" * 60)
print(f"Initial: {rabbits.value:.0f} rabbits, {foxes.value:.0f} foxes")
print("\n📋 Competing Ecological Theories:")
print(" 1. Lotka-Volterra (classic predator-prey cycles)")
print(" 2. Logistic Prey (carrying capacity limits prey)")
print(" 3. Ratio Model (maintains fixed predator:prey ratio)")
print("\n" + "-" * 60 + "\n")
# Run simulation
executive.run(steps=100)
print("\n" + "=" * 60)
print("Simulation Complete!")
print(f"Final: {rabbits.value:.0f} rabbits, {foxes.value:.0f} foxes")
if stability_gov.interventions:
print(f"Ecosystem interventions: {len(stability_gov.interventions)}")
print("=" * 60)
Step 5: Analyze Coupled Dynamics
def analyze_coupled_system() -> None:
"""Analyze the coupled predator-prey dynamics."""
if rabbits.memory is None or foxes.memory is None:
return
rabbits_history = [r.value for _, r, _ in rabbits.memory.records() if r is not None]
foxes_history = [r.value for _, r, _ in foxes.memory.records() if r is not None]
print("\n📊 Ecosystem Dynamics Analysis")
print("-" * 40)
# 1. Basic statistics
print("\n🐇 Rabbits:")
print(f" Initial: {rabbits_history[0]:.0f}")
print(f" Final: {rabbits_history[-1]:.0f}")
print(f" Mean: {np.mean(rabbits_history):.0f}")
print(f" Std Dev: {np.std(rabbits_history):.0f}")
print(f" Min: {np.min(rabbits_history):.0f}")
print(f" Max: {np.max(rabbits_history):.0f}")
print("\n🦊 Foxes:")
print(f" Initial: {foxes_history[0]:.0f}")
print(f" Final: {foxes_history[-1]:.0f}")
print(f" Mean: {np.mean(foxes_history):.0f}")
print(f" Std Dev: {np.std(foxes_history):.0f}")
print(f" Min: {np.min(foxes_history):.0f}")
print(f" Max: {np.max(foxes_history):.0f}")
# 2. Correlation analysis
correlation = np.corrcoef(rabbits_history, foxes_history)[0, 1]
print(f"\n📈 Rabbit-Fox Correlation: {correlation:.3f}")
if correlation > 0.3:
print(" → Populations move together (positive correlation)")
elif correlation < -0.3:
print(
" → Populations move opposite (negative correlation - "
"classic predator-prey)"
)
else:
print(" → Weak correlation (other dynamics at play)")
# 3. Cycle detection
rabbit_peaks, _ = find_peaks(rabbits_history, height=np.mean(rabbits_history))
fox_peaks, _ = find_peaks(foxes_history, height=np.mean(foxes_history))
print("\n🔄 Cycles detected:")
print(f" Rabbit peaks: {len(rabbit_peaks)}")
print(f" Fox peaks: {len(fox_peaks)}")
if len(rabbit_peaks) > 1:
avg_cycle = np.mean(np.diff(rabbit_peaks))
print(f" Avg rabbit cycle length: {avg_cycle:.0f} steps")
# 4. Predator-prey ratio
ratios = [r / f if f > 0 else 0 for r, f in zip(rabbits_history, foxes_history)]
print("\n⚖️ Rabbit:Fox Ratio:")
print(f" Mean: {np.mean(ratios):.2f}")
print(f" Std: {np.std(ratios):.2f}")
print(f" Min: {np.min(ratios):.2f}")
print(f" Max: {np.max(ratios):.2f}")
# 5. Extinction risk
rabbit_extinction_risk = sum(1 for r in rabbits_history if r < 10) / len(
rabbits_history
)
fox_extinction_risk = sum(1 for f in foxes_history if f < 10) / len(foxes_history)
print("\n⚠️ Extinction Risk (% steps below 10):")
print(f" Rabbits: {rabbit_extinction_risk*100:.1f}%")
print(f" Foxes: {fox_extinction_risk*100:.1f}%")
# Run analysis
analyze_coupled_system()
Step 6: Visualization of Coupled Dynamics
def plot_coupled_dynamics(stability_gov: EcosystemStabilityGovernance) -> None:
"""Visualize the coupled predator-prey dynamics."""
if rabbits.memory is None or foxes.memory is None:
return
rabbits_history = [r.value for _, r, _ in rabbits.memory.records() if r is not None]
foxes_history = [r.value for _, r, _ in foxes.memory.records() if r is not None]
steps = list(range(len(rabbits_history)))
# Create figure with subplots
fig = plt.figure(figsize=(12, 10))
gs = GridSpec(3, 2, figure=fig, height_ratios=[2, 1, 1])
# Plot 1: Populations over time
ax1 = fig.add_subplot(gs[0, :])
ax1.plot(steps, rabbits_history, "b-", linewidth=2, label="Rabbits (Prey)")
ax1.plot(steps, foxes_history, "r-", linewidth=2, label="Foxes (Predator)")
# Highlight stability interventions
for intervention in stability_gov.interventions:
step = intervention["step"]
if step < len(steps):
ax1.axvline(x=step, color="orange", linestyle="--", alpha=0.7, linewidth=1)
ax1.text(
step,
ax1.get_ylim()[1] * 0.9,
"⚡",
fontsize=12,
ha="center",
color="orange",
)
ax1.set_xlabel("Step")
ax1.set_ylabel("Population")
ax1.set_title("Predator-Prey Population Dynamics")
ax1.legend(loc="upper right")
ax1.grid(True, alpha=0.3)
# Plot 2: Phase portrait (Rabbits vs Foxes)
ax2 = fig.add_subplot(gs[1, 0])
ax2.plot(rabbits_history, foxes_history, "g-", alpha=0.7, linewidth=1)
ax2.scatter(
rabbits_history[0],
foxes_history[0],
color="green",
s=100,
label="Start",
zorder=5,
)
ax2.scatter(
rabbits_history[-1],
foxes_history[-1],
color="red",
s=100,
label="End",
zorder=5,
)
ax2.set_xlabel("Rabbits")
ax2.set_ylabel("Foxes")
ax2.set_title("Phase Portrait (State Space)")
ax2.legend()
ax2.grid(True, alpha=0.3)
# Plot 3: Predator-Prey Ratio
ax3 = fig.add_subplot(gs[1, 1])
ratios = [r / f if f > 0 else 0 for r, f in zip(rabbits_history, foxes_history)]
ax3.plot(steps, ratios, "purple", linewidth=2)
ax3.axhline(
y=np.mean(ratios),
color="purple",
linestyle="--",
alpha=0.5,
label=f"Mean: {np.mean(ratios):.2f}",
)
ax3.set_xlabel("Step")
ax3.set_ylabel("Rabbit:Fox Ratio")
ax3.set_title("Predator-Prey Ratio")
ax3.legend()
ax3.grid(True, alpha=0.3)
# Plot 4: Cross-correlation
ax4 = fig.add_subplot(gs[2, :])
# Calculate cross-correlation
max_lag = min(20, len(rabbits_history) // 4)
correlation = np.correlate(
rabbits_history - np.mean(rabbits_history),
foxes_history - np.mean(foxes_history),
mode="full",
)
correlation = correlation / (
len(rabbits_history) * np.std(rabbits_history) * np.std(foxes_history)
)
lags = np.arange(-max_lag, max_lag + 1)
center = len(correlation) // 2
correlation_trimmed = correlation[center - max_lag : center + max_lag + 1]
ax4.bar(lags, correlation_trimmed, alpha=0.7)
ax4.axhline(y=0, color="black", linestyle="-", linewidth=0.5)
ax4.axvline(x=0, color="red", linestyle="--", alpha=0.5, label="No lag")
ax4.set_xlabel("Lag (Foxes relative to Rabbits)")
ax4.set_ylabel("Correlation")
ax4.set_title("Cross-Correlation: Rabbits vs Foxes")
ax4.legend()
ax4.grid(True, alpha=0.3)
plt.tight_layout()
plt.savefig("predator_prey_dynamics.png", dpi=150)
plt.show()
# Uncomment to visualize
#plot_coupled_dynamics(stability_gov)
Complete Script
Here's the complete, runnable example:
#!/usr/bin/env python3
"""Multiple Variables - Predator-Prey Ecosystem with Competing Models"""
from typing import Any
import matplotlib.pyplot as plt
import numpy as np
from matplotlib.gridspec import GridSpec
from scipy.signal import find_peaks
from procela import (
Executive,
InvariantPhase,
InvariantViolation,
Key,
Mechanism,
RangeDomain,
SystemInvariant,
Variable,
VariableRecord,
VariableSnapshot,
WeightedConfidencePolicy,
)
rng = np.random.default_rng(42)
INITIAL_SOURCE = Key()
# ========================================================
# Prey population (Rabbits) - range 0 to 1000
rabbits = Variable(
name="Rabbits", domain=RangeDomain(0, 1000), policy=WeightedConfidencePolicy()
)
# Predator population (Foxes) - range 0 to 500
foxes = Variable(
name="Foxes", domain=RangeDomain(0, 500), policy=WeightedConfidencePolicy()
)
# Initialize populations
rabbits.init(VariableRecord(100.0, confidence=1.0, source=INITIAL_SOURCE))
foxes.init(VariableRecord(20.0, confidence=1.0, source=INITIAL_SOURCE))
print(f"Initial ecosystem: {rabbits.value:.0f} rabbits, {foxes.value:.0f} foxes")
# ===========================================================
class LotkaVolterraMechanism(Mechanism):
"""
Lotka volterra mechanism.
Classic predator-prey model:
- Prey growth: dr/dt = α*r - β*r*f
- Predator growth: df/dt = δ*r*f - γ*f
where:
α = prey birth rate
β = predation rate
δ = predator efficiency (how well predators convert prey to offspring)
γ = predator death rate
"""
def __init__(
self,
alpha: float = 0.08,
beta: float = 0.02,
delta: float = 0.01,
gamma: float = 0.06,
) -> None:
"""Lotka volterra mechanism constructor."""
super().__init__(reads=[rabbits, foxes], writes=[rabbits, foxes])
self.alpha = alpha # Prey birth rate
self.beta = beta # Predation rate
self.delta = delta # Predator efficiency
self.gamma = gamma # Predator death rate
def transform(self) -> None:
"""Transform method."""
# Read current populations
r = self.reads()[0].value # Rabbits
f = self.reads()[1].value # Foxes
# Calculate changes
dr_dt = r * (self.alpha - self.beta * f)
df_dt = f * (self.delta * r - self.gamma)
# Discrete time update (dt = 1)
new_rabbits = max(0, r + dr_dt)
new_foxes = max(0, f + df_dt)
# Add stochastic noise (environmental variability)
new_rabbits += rng.normal(0, new_rabbits * 0.05)
new_foxes += rng.normal(0, new_foxes * 0.05)
# Confidence decreases with extreme values
rabbit_confidence = 0.9 if 10 < new_rabbits < 500 else 0.6
fox_confidence = 0.9 if 5 < new_foxes < 200 else 0.6
# Propose hypotheses for both variables
self.writes()[0].add_hypothesis(
VariableRecord(
value=new_rabbits,
confidence=rabbit_confidence,
source=self.key(),
metadata={"model": "classic", "alpha": self.alpha},
)
)
self.writes()[1].add_hypothesis(
VariableRecord(
value=new_foxes,
confidence=fox_confidence,
source=self.key(),
metadata={"model": "classic", "gamma": self.gamma},
)
)
print(
f" 🦊 LV Model: R:{new_rabbits:.0f} (conf:{rabbit_confidence:.2f}), "
f"F:{new_foxes:.0f} (conf:{fox_confidence:.2f})"
)
# ============================================================
class LogisticPreyMechanism(Mechanism):
"""
Alternative model.
Prey growth is logistic (carrying capacity limited) rather than exponential.
"""
def __init__(
self,
carrying_capacity: float = 800,
predation_rate: float = 0.03,
predator_efficiency: float = 0.008,
predator_death: float = 0.07,
) -> None:
"""Logistic prey mechanism constructor."""
super().__init__(reads=[rabbits, foxes], writes=[rabbits, foxes])
self.K = carrying_capacity
self.beta = predation_rate
self.delta = predator_efficiency
self.gamma = predator_death
def transform(self) -> None:
"""Transform method."""
r = self.reads()[0].value
f = self.reads()[1].value
# Logistic prey growth: r * (1 - r/K)
prey_growth = r * (1 - r / self.K)
prey_death = self.beta * r * f
new_rabbits = max(0, r + prey_growth - prey_death)
# Predator growth (same as LV)
predator_growth = self.delta * r * f
predator_death = self.gamma * f
new_foxes = max(0, f + predator_growth - predator_death)
# Add noise
new_rabbits += rng.normal(0, new_rabbits * 0.03)
new_foxes += rng.normal(0, new_foxes * 0.03)
# Higher confidence when near equilibrium
equilibrium_r = self.K * (1 - self.gamma / (self.delta * self.K))
r_distance = abs(new_rabbits - equilibrium_r) / self.K
rabbit_confidence = max(0.5, 0.9 - r_distance)
self.writes()[0].add_hypothesis(
VariableRecord(
new_rabbits,
confidence=rabbit_confidence,
source=self.key(),
metadata={"model": "logistic"},
)
)
self.writes()[1].add_hypothesis(
VariableRecord(
new_foxes,
confidence=0.8,
source=self.key(),
metadata={"model": "logistic"},
)
)
print(
f" 🌿 Logistic Model: R:{new_rabbits:.0f} (conf:{rabbit_confidence:.2f}), "
f"F:{new_foxes:.0f}"
)
# =========================================================
class RatioMechanism(Mechanism):
"""
Simple model based on predator-prey ratio.
Assumes populations tend toward a fixed ratio.
"""
def __init__(self, target_ratio: float = 5.0, adjustment_rate: float = 0.1) -> None:
"""Ratio mechanism constructor."""
super().__init__(reads=[rabbits, foxes], writes=[rabbits, foxes])
self.target_ratio = target_ratio # Target rabbits:foxes ratio
self.rate = adjustment_rate
def transform(self) -> None:
"""Transform method."""
r = self.reads()[0].value
f = self.reads()[1].value
if f == 0:
new_foxes = 1.0 # Avoid division by zero
else:
current_ratio = r / f
# Adjust toward target ratio
ratio_error = current_ratio - self.target_ratio
adjustment = self.rate * ratio_error
# Apply adjustment
new_rabbits = max(0, r - adjustment * r)
new_foxes = max(0, f + adjustment * f)
# Simple model has lower confidence
confidence = 0.65
self.writes()[0].add_hypothesis(
VariableRecord(new_rabbits, confidence=confidence, source=self.key())
)
self.writes()[1].add_hypothesis(
VariableRecord(new_foxes, confidence=confidence, source=self.key())
)
print(
f" ⚖️ Ratio Model: R:{new_rabbits:.0f}, "
f"F:{new_foxes:.0f} (conf:{confidence:.2f})"
)
# ===============================================================
class EcosystemStabilityGovernance(SystemInvariant):
"""Monitor ecosystem health and prevents extinction."""
def __init__(
self, rabbits_var: Variable, foxes_var: Variable, min_population: float = 10
) -> None:
"""Ecosystem stability governance constructor."""
self.rabbits = rabbits_var
self.foxes = foxes_var
self.min_population = min_population
self.interventions: list[dict[str, Any]] = []
def check(snapshot: VariableSnapshot) -> bool:
"""Check ecosystem health after each step."""
r = self.rabbits.value
f = self.foxes.value
if r < self.min_population or f < self.min_population:
print(f"\n 🚨 ECOSYSTEM CRISIS at step {snapshot.step}!")
print(f" Population too low: R={r:.0f}, F={f:.0f}")
# Intervention: Restore populations
if r < self.min_population:
print(
" Intervention: Restoring rabbit population "
f"to {self.min_population}"
)
if f < self.min_population:
print(
" Intervention: Restoring fox population "
f"to {self.min_population}"
)
self.interventions.append(
{"step": snapshot.step, "rabbits_before": r, "foxes_before": f}
)
return False
return True
def handle(invariant: InvariantViolation, snapshot: VariableSnapshot) -> None:
# Nothing to handle
pass
super().__init__(
name="EcosystemStability",
condition=check,
on_violation=handle,
phase=InvariantPhase.POST,
)
# We'll integrate this into the executive step
# ===========================================================
# Create all mechanisms
mechanisms = [
LotkaVolterraMechanism(alpha=0.08, beta=0.02, delta=0.01, gamma=0.06),
LogisticPreyMechanism(carrying_capacity=800, predation_rate=0.025),
RatioMechanism(target_ratio=4.0, adjustment_rate=0.08),
]
# Create stability governance
stability_gov = EcosystemStabilityGovernance(rabbits, foxes, min_population=10)
# Create executive
executive = Executive(mechanisms=mechanisms)
executive.add_invariant(stability_gov)
print("\n" + "=" * 60)
print("Predator-Prey Ecosystem Simulation")
print("=" * 60)
print(f"Initial: {rabbits.value:.0f} rabbits, {foxes.value:.0f} foxes")
print("\n📋 Competing Ecological Theories:")
print(" 1. Lotka-Volterra (classic predator-prey cycles)")
print(" 2. Logistic Prey (carrying capacity limits prey)")
print(" 3. Ratio Model (maintains fixed predator:prey ratio)")
print("\n" + "-" * 60 + "\n")
# Run simulation
executive.run(steps=100)
print("\n" + "=" * 60)
print("Simulation Complete!")
print(f"Final: {rabbits.value:.0f} rabbits, {foxes.value:.0f} foxes")
if stability_gov.interventions:
print(f"Ecosystem interventions: {len(stability_gov.interventions)}")
print("=" * 60)
# ===========================================================
def analyze_coupled_system() -> None:
"""Analyze the coupled predator-prey dynamics."""
if rabbits.memory is None or foxes.memory is None:
return
rabbits_history = [r.value for _, r, _ in rabbits.memory.records() if r is not None]
foxes_history = [r.value for _, r, _ in foxes.memory.records() if r is not None]
print("\n📊 Ecosystem Dynamics Analysis")
print("-" * 40)
# 1. Basic statistics
print("\n🐇 Rabbits:")
print(f" Initial: {rabbits_history[0]:.0f}")
print(f" Final: {rabbits_history[-1]:.0f}")
print(f" Mean: {np.mean(rabbits_history):.0f}")
print(f" Std Dev: {np.std(rabbits_history):.0f}")
print(f" Min: {np.min(rabbits_history):.0f}")
print(f" Max: {np.max(rabbits_history):.0f}")
print("\n🦊 Foxes:")
print(f" Initial: {foxes_history[0]:.0f}")
print(f" Final: {foxes_history[-1]:.0f}")
print(f" Mean: {np.mean(foxes_history):.0f}")
print(f" Std Dev: {np.std(foxes_history):.0f}")
print(f" Min: {np.min(foxes_history):.0f}")
print(f" Max: {np.max(foxes_history):.0f}")
# 2. Correlation analysis
correlation = np.corrcoef(rabbits_history, foxes_history)[0, 1]
print(f"\n📈 Rabbit-Fox Correlation: {correlation:.3f}")
if correlation > 0.3:
print(" → Populations move together (positive correlation)")
elif correlation < -0.3:
print(
" → Populations move opposite (negative correlation - "
"classic predator-prey)"
)
else:
print(" → Weak correlation (other dynamics at play)")
# 3. Cycle detection
rabbit_peaks, _ = find_peaks(rabbits_history, height=np.mean(rabbits_history))
fox_peaks, _ = find_peaks(foxes_history, height=np.mean(foxes_history))
print("\n🔄 Cycles detected:")
print(f" Rabbit peaks: {len(rabbit_peaks)}")
print(f" Fox peaks: {len(fox_peaks)}")
if len(rabbit_peaks) > 1:
avg_cycle = np.mean(np.diff(rabbit_peaks))
print(f" Avg rabbit cycle length: {avg_cycle:.0f} steps")
# 4. Predator-prey ratio
ratios = [r / f if f > 0 else 0 for r, f in zip(rabbits_history, foxes_history)]
print("\n⚖️ Rabbit:Fox Ratio:")
print(f" Mean: {np.mean(ratios):.2f}")
print(f" Std: {np.std(ratios):.2f}")
print(f" Min: {np.min(ratios):.2f}")
print(f" Max: {np.max(ratios):.2f}")
# 5. Extinction risk
rabbit_extinction_risk = sum(1 for r in rabbits_history if r < 10) / len(
rabbits_history
)
fox_extinction_risk = sum(1 for f in foxes_history if f < 10) / len(foxes_history)
print("\n⚠️ Extinction Risk (% steps below 10):")
print(f" Rabbits: {rabbit_extinction_risk*100:.1f}%")
print(f" Foxes: {fox_extinction_risk*100:.1f}%")
# Run analysis
analyze_coupled_system()
# ===============================================================
def plot_coupled_dynamics(stability_gov: EcosystemStabilityGovernance) -> None:
"""Visualize the coupled predator-prey dynamics."""
if rabbits.memory is None or foxes.memory is None:
return
rabbits_history = [r.value for _, r, _ in rabbits.memory.records() if r is not None]
foxes_history = [r.value for _, r, _ in foxes.memory.records() if r is not None]
steps = list(range(len(rabbits_history)))
# Create figure with subplots
fig = plt.figure(figsize=(12, 10))
gs = GridSpec(3, 2, figure=fig, height_ratios=[2, 1, 1])
# Plot 1: Populations over time
ax1 = fig.add_subplot(gs[0, :])
ax1.plot(steps, rabbits_history, "b-", linewidth=2, label="Rabbits (Prey)")
ax1.plot(steps, foxes_history, "r-", linewidth=2, label="Foxes (Predator)")
# Highlight stability interventions
for intervention in stability_gov.interventions:
step = intervention["step"]
if step < len(steps):
ax1.axvline(x=step, color="orange", linestyle="--", alpha=0.7, linewidth=1)
ax1.text(
step,
ax1.get_ylim()[1] * 0.9,
"⚡",
fontsize=12,
ha="center",
color="orange",
)
ax1.set_xlabel("Step")
ax1.set_ylabel("Population")
ax1.set_title("Predator-Prey Population Dynamics")
ax1.legend(loc="upper right")
ax1.grid(True, alpha=0.3)
# Plot 2: Phase portrait (Rabbits vs Foxes)
ax2 = fig.add_subplot(gs[1, 0])
ax2.plot(rabbits_history, foxes_history, "g-", alpha=0.7, linewidth=1)
ax2.scatter(
rabbits_history[0],
foxes_history[0],
color="green",
s=100,
label="Start",
zorder=5,
)
ax2.scatter(
rabbits_history[-1],
foxes_history[-1],
color="red",
s=100,
label="End",
zorder=5,
)
ax2.set_xlabel("Rabbits")
ax2.set_ylabel("Foxes")
ax2.set_title("Phase Portrait (State Space)")
ax2.legend()
ax2.grid(True, alpha=0.3)
# Plot 3: Predator-Prey Ratio
ax3 = fig.add_subplot(gs[1, 1])
ratios = [r / f if f > 0 else 0 for r, f in zip(rabbits_history, foxes_history)]
ax3.plot(steps, ratios, "purple", linewidth=2)
ax3.axhline(
y=np.mean(ratios),
color="purple",
linestyle="--",
alpha=0.5,
label=f"Mean: {np.mean(ratios):.2f}",
)
ax3.set_xlabel("Step")
ax3.set_ylabel("Rabbit:Fox Ratio")
ax3.set_title("Predator-Prey Ratio")
ax3.legend()
ax3.grid(True, alpha=0.3)
# Plot 4: Cross-correlation
ax4 = fig.add_subplot(gs[2, :])
# Calculate cross-correlation
max_lag = min(20, len(rabbits_history) // 4)
correlation = np.correlate(
rabbits_history - np.mean(rabbits_history),
foxes_history - np.mean(foxes_history),
mode="full",
)
correlation = correlation / (
len(rabbits_history) * np.std(rabbits_history) * np.std(foxes_history)
)
lags = np.arange(-max_lag, max_lag + 1)
center = len(correlation) // 2
correlation_trimmed = correlation[center - max_lag : center + max_lag + 1]
ax4.bar(lags, correlation_trimmed, alpha=0.7)
ax4.axhline(y=0, color="black", linestyle="-", linewidth=0.5)
ax4.axvline(x=0, color="red", linestyle="--", alpha=0.5, label="No lag")
ax4.set_xlabel("Lag (Foxes relative to Rabbits)")
ax4.set_ylabel("Correlation")
ax4.set_title("Cross-Correlation: Rabbits vs Foxes")
ax4.legend()
ax4.grid(True, alpha=0.3)
plt.tight_layout()
plt.savefig("predator_prey_dynamics.png", dpi=150)
plt.show()
# Uncomment to visualize
#plot_coupled_dynamics(stability_gov)
Expected Output
Initial ecosystem: 100 rabbits, 20 foxes
============================================================
Predator-Prey Ecosystem Simulation
============================================================
Initial: 100 rabbits, 20 foxes
📋 Competing Ecological Theories:
1. Lotka-Volterra (classic predator-prey cycles)
2. Logistic Prey (carrying capacity limits prey)
3. Ratio Model (maintains fixed predator:prey ratio)
------------------------------------------------------------
🦊 LV Model: R:69 (conf:0.90), F:37 (conf:0.90)
🌿 Logistic Model: R:141 (conf:0.50), F:36
⚖️ Ratio Model: R:92, F:22 (conf:0.65)
🦊 LV Model: R:37 (conf:0.90), F:56 (conf:0.90)
🌿 Logistic Model: R:102 (conf:0.50), F:54
⚖️ Ratio Model: R:102, F:29 (conf:0.65)
🦊 LV Model: R:9 (conf:0.60), F:77 (conf:0.90)
🌿 Logistic Model: R:53 (conf:0.50), F:74
⚖️ Ratio Model: R:88, F:38 (conf:0.65)
🦊 LV Model: R:0 (conf:0.60), F:100 (conf:0.90)
🌿 Logistic Model: R:16 (conf:0.50), F:85
⚖️ Ratio Model: R:64, F:49 (conf:0.65)
🦊 LV Model: R:0 (conf:0.60), F:94 (conf:0.90)
🌿 Logistic Model: R:0 (conf:0.50), F:93
⚖️ Ratio Model: R:36, F:57 (conf:0.65)
🦊 LV Model: R:0 (conf:0.60), F:87 (conf:0.90)
🌿 Logistic Model: R:0 (conf:0.50), F:86
⚖️ Ratio Model: R:18, F:58 (conf:0.65)
🚨 ECOSYSTEM CRISIS at step 5!
Population too low: R=7, F=79
Intervention: Restoring rabbit population to 10
🦊 LV Model: R:0 (conf:0.60), F:78 (conf:0.90)
🌿 Logistic Model: R:0 (conf:0.50), F:78
⚖️ Ratio Model: R:9, F:54 (conf:0.65)
🚨 ECOSYSTEM CRISIS at step 6!
Population too low: R=3, F=71
Intervention: Restoring rabbit population to 10
🦊 LV Model: R:0 (conf:0.60), F:71 (conf:0.90)
🌿 Logistic Model: R:1 (conf:0.50), F:67
⚖️ Ratio Model: R:4, F:49 (conf:0.65)
🚨 ECOSYSTEM CRISIS at step 7!
Population too low: R=2, F=64
Intervention: Restoring rabbit population to 10
🦊 LV Model: R:0 (conf:0.60), F:58 (conf:0.90)
🌿 Logistic Model: R:1 (conf:0.50), F:62
⚖️ Ratio Model: R:2, F:43 (conf:0.65)
[Simulation runs...]
🦊 LV Model: R:14 (conf:0.90), F:17 (conf:0.90)
🌿 Logistic Model: R:29 (conf:0.50), F:16
⚖️ Ratio Model: R:22, F:12 (conf:0.65)
============================================================
Simulation Complete!
Final: 20 rabbits, 15 foxes
Ecosystem interventions: 63
============================================================
📊 Ecosystem Dynamics Analysis
----------------------------------------
🐇 Rabbits:
Initial: 100
Final: 20
Mean: 18
Std Dev: 20
Min: 0
Max: 100
🦊 Foxes:
Initial: 20
Final: 15
Mean: 26
Std Dev: 19
Min: 4
Max: 84
📈 Rabbit-Fox Correlation: -0.003
→ Weak correlation (other dynamics at play)
🔄 Cycles detected:
Rabbit peaks: 2
Fox peaks: 3
Avg rabbit cycle length: 34 steps
⚖️ Rabbit:Fox Ratio:
Mean: 1.18
Std: 1.45
Min: 0.01
Max: 5.33
⚠️ Extinction Risk (% steps below 10):
Rabbits: 49.5%
Foxes: 20.8%
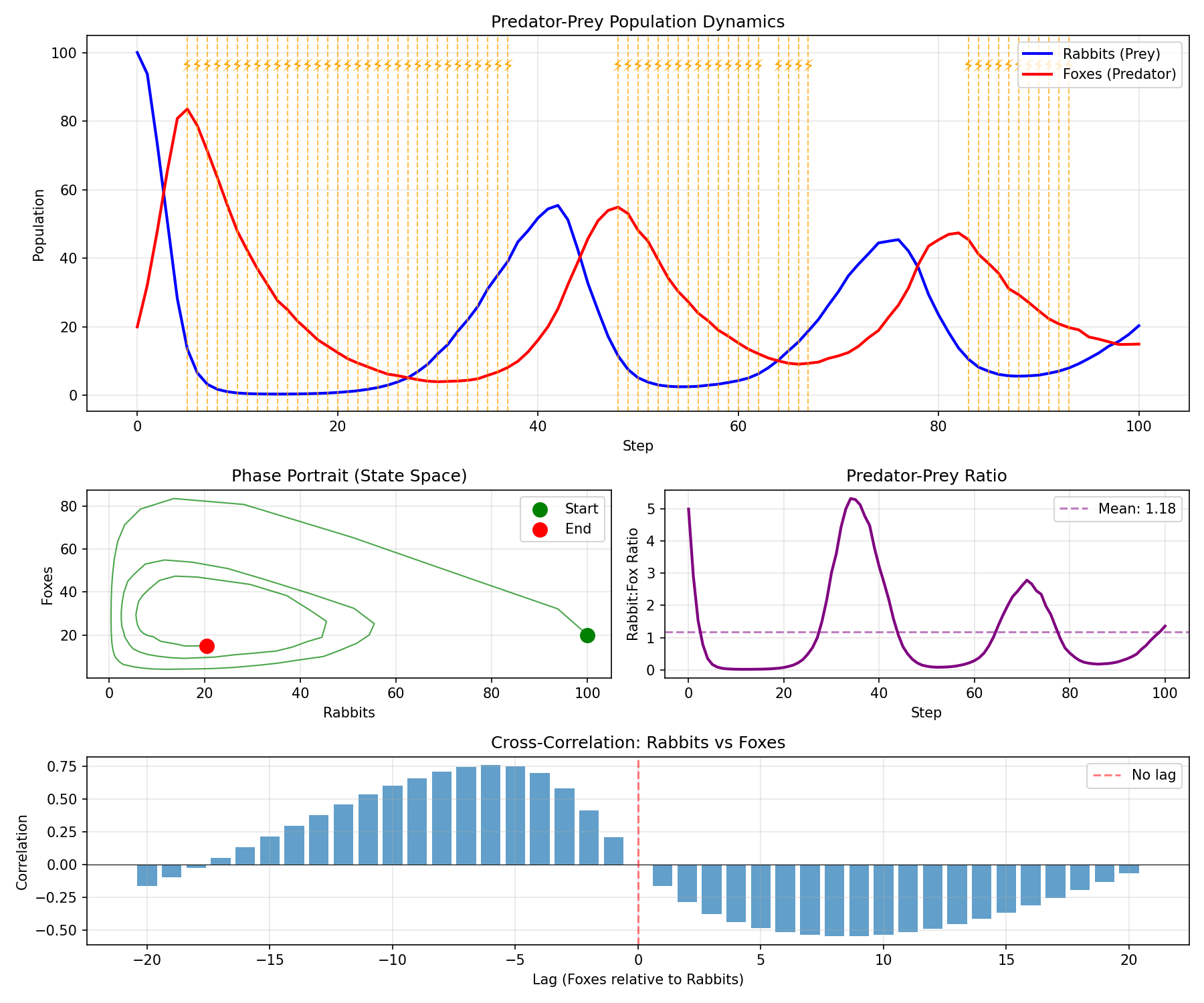
Key Takeaways
- Variables can be coupled - Mechanisms read from multiple variables and write to multiple variables
- Competing theories - Different ecological models propose different dynamics
- Emergent behavior - The resolution process determines which model "wins" at each step
- System-level governance - Monitor and intervene at ecosystem level, not just individual variables
- Phase portraits - Visualizing multiple variables together reveals system dynamics
Exercises
Try modifying the example to explore:
-
Add a third variable - Introduce grass (food for rabbits) to create a three-level food chain
-
Seasonal forcing - Make birth rates vary sinusoidally over time (seasons)
-
Spatial patches - Create multiple coupled rabbit-fox pairs with migration between patches
-
Harvesting mechanism - Add human intervention that removes a percentage of both populations
-
Alternative governance - Implement a governance that maintains biodiversity by preventing any population from dominating
Next Steps
- Learn about Epistemic Signals for advanced monitoring
- Explore Intermediate Governance patterns
- See the AMR Case Study with multiple interacting variables
- Study Advanced Multi-Variable Systems for optimization
Troubleshooting
| Issue | Solution |
|---|---|
| Populations go negative | Check max(0, value) in mechanism transforms |
| One variable dominates | Adjust confidence scores or add governance to balance |
| Extinction occurs frequently | Lower min_population threshold or adjust growth rates |
| Variables not updating | Ensure mechanisms include all variables in writes list |